Microorganism confirmation and identification are critical in a number of industries, including pharma, cosmetics, and food. In the food industry, pressures from regulatory bodies and customers alike are continually tightening, making accurate, rapid microbiology testing crucial for manufacturers to make rapid decisions in regard to quality and safety, for example, to detect possible food spoilage organisms or foodborne pathogens, or monitor environmental contaminants or technological microflora.
A wide variety of technologies are used to detect and identify contaminants and technological organisms in food, from traditional culture-based methods and molecular testing using polymerase chain reaction to increasingly sophisticated analytical technologies such as mass spectrometry (MS). The ability to identify bacteria, yeast, and mold at the genus or species level enables a deeper level of screening. Researchers and test manufacturers are always searching for more advanced methods for faster tests with higher accuracy and sensitivity.
The Benefits of MS
Food safety authorities have fully accepted the AOAC- and ISO 16140-approved rapid testing methods, and their adoption has been acknowledged in regional regulations. The development of new technologies, such as matrix-assisted laser desorption/ionization (MALDI) time-of-flight (TOF) MS, and sequencing offers faster and more accurate confirmation and identification of microorganisms to the food microbiology industry. Time is often the most critical factor in food microbiology laboratories, so MALDI-TOF is becoming increasingly valuable in routine testing.
Comparative studies have demonstrated differences in the percentages of correctly identified bacteria and fungi isolates using MALDI-TOF MS (>99%) and traditional identification panels (94–97%), with speed and reagent costs also representing key benefits of MALDI-TOF MS.[1] MALDI-TOF MS will rapidly establish a robust position for surveillance and monitoring of microorganisms in the complex interwoven network of origin, producers, traders, and consumers.[2] For instance, in 2016, more than 1,200 Listeria spp. strains—in the context of national surveillance in France—were analyzed using MALDI-TOF MS. All isolates were correctly identified by MALDI-TOF MS at the species level, as confirmed by whole-genome sequencing.[3]
MALDI-TOF MS in Practice
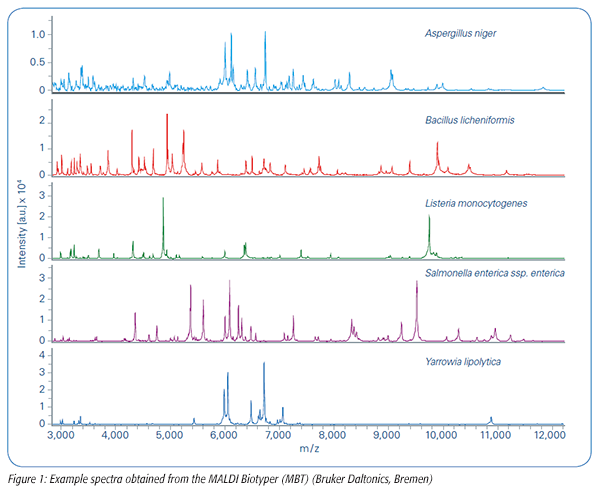 In January 2017, the Microbiology Food Laboratory at Q Laboratories in Cincinnati, OH, added Bruker’s MALDI-TOF MS system, the MALDI Biotyper (MBT), to its toolbox of technologies for routine food microbial identification and research and development work. This technology determines a unique protein fingerprint of a microorganism, which is used to reliably identify a particular species by matching the fingerprint with thousands of reference spectra from a comprehensive library (Figure 1). This tool is complemented by existing techniques in the toolbox, such as sequence-based identification with rDNA sequencing.
In January 2017, the Microbiology Food Laboratory at Q Laboratories in Cincinnati, OH, added Bruker’s MALDI-TOF MS system, the MALDI Biotyper (MBT), to its toolbox of technologies for routine food microbial identification and research and development work. This technology determines a unique protein fingerprint of a microorganism, which is used to reliably identify a particular species by matching the fingerprint with thousands of reference spectra from a comprehensive library (Figure 1). This tool is complemented by existing techniques in the toolbox, such as sequence-based identification with rDNA sequencing.
Routine Testing
In the routine laboratory, the incorporation of MS technology into the toolbox enables the laboratory to offer flexible options for turnaround time and cost to fit in with the client’s needs. This is particularly valuable as it creates a completely customizable service and provides the option of same-day results for rapid pathogen identification, for example. This was not available with previous methods. The confidence in results with the MALDI-TOF system and increased sensitivity means that fewer re-tests are required, so when an identification is provided for the first time, the client can make decisions immediately.
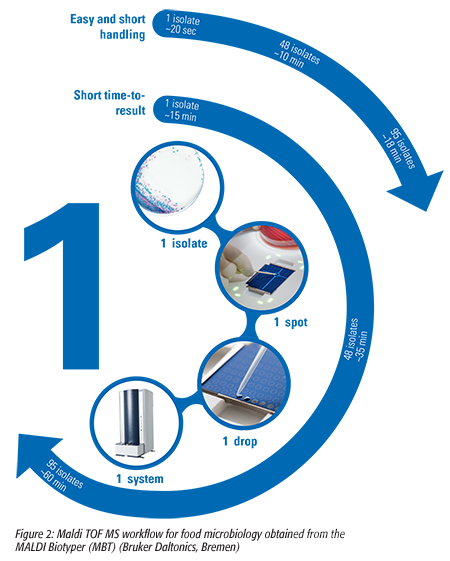 The Food Microbiology Laboratory now analyzes samples with the MBT in 15 minutes to less than 1 hour to either confirm presumptive results of a detection or enumeration method or to identify an organism from a plate. This encompasses 20 seconds of handling time, and restricted consumables are required for more than 95 percent of the microbial isolates, providing a cost-effective microbial confirmation and identification solution (Figure 2). There is no need to know the type of microorganism prior to analysis; bacteria, yeast, or mold samples are all analyzed together, following the same workflow.
The Food Microbiology Laboratory now analyzes samples with the MBT in 15 minutes to less than 1 hour to either confirm presumptive results of a detection or enumeration method or to identify an organism from a plate. This encompasses 20 seconds of handling time, and restricted consumables are required for more than 95 percent of the microbial isolates, providing a cost-effective microbial confirmation and identification solution (Figure 2). There is no need to know the type of microorganism prior to analysis; bacteria, yeast, or mold samples are all analyzed together, following the same workflow.
Validating New Technologies
The Q Laboratories R&D laboratory tests and validates technologies and assays for food microbiology testing. Technology companies and diagnostic partners approach Q Laboratories with a new technology or assay for method validation, for example, for testing Salmonella, Listeria, or Legionella, and to request a proposal for the work. The R&D laboratory establishes a study design with the client and the method validation organization (i.e., AOAC INTERNATIONAL in the U.S.). The study should fulfill the AOAC Appendix J guidelines requirements.[4]
MicroVal and AFNOR Certification are the certification bodies in Europe, and method performance is assessed using the ISO 16140 standard.[5, 6] The "Expert Laboratory" (in this case, Q Laboratories) is the party that prepares the protocol, in collaboration with the technology provider, and submits the protocol to the certification body for the technical reviewers to approve. Q Laboratories is the only North American laboratory to be an Expert Laboratory for all three of the major certification bodies: AOAC, MicroVal, and AFNOR Certification. This enables the design of combined AOAC/ISO validation studies and fulfillment of all client requirements.
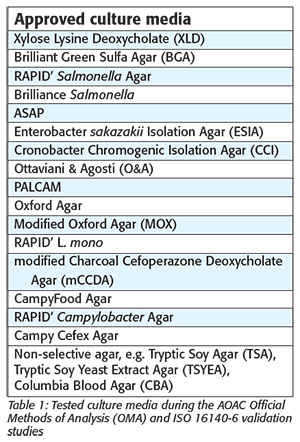 Q Laboratories conducted AOAC Official Methods of Analysis (OMA) studies and ISO 16140 validation studies with MicroVal certification body for the Bruker MBT. By combining existing guidelines—such as the AOAC Appendix J guidelines and the new ISO 16140 part 6 standard[6]—an optimal study design was created to validate the use of the MBT for food microbiology analysis. These studies were the first of their kind, as the designs for validating a confirmation and identification method had not caught up with the newest technology. Fifty percent of the data were produced in the U.S. and the other half in Europe, by collaborating with a second Expert Laboratory, ADRIA (France). This enabled the laboratories to test the relevant biodiversity of strains from various origins to verify the reliability of the MBT. In addition, four collaborative studies involving laboratories from two different regions were run to assess the reproducibility and robustness of the MBT. Various selective culture media were tested (Table 1), as well as two instruments, the MBT and MBT-smart.
Q Laboratories conducted AOAC Official Methods of Analysis (OMA) studies and ISO 16140 validation studies with MicroVal certification body for the Bruker MBT. By combining existing guidelines—such as the AOAC Appendix J guidelines and the new ISO 16140 part 6 standard[6]—an optimal study design was created to validate the use of the MBT for food microbiology analysis. These studies were the first of their kind, as the designs for validating a confirmation and identification method had not caught up with the newest technology. Fifty percent of the data were produced in the U.S. and the other half in Europe, by collaborating with a second Expert Laboratory, ADRIA (France). This enabled the laboratories to test the relevant biodiversity of strains from various origins to verify the reliability of the MBT. In addition, four collaborative studies involving laboratories from two different regions were run to assess the reproducibility and robustness of the MBT. Various selective culture media were tested (Table 1), as well as two instruments, the MBT and MBT-smart.
Q Laboratories served as the lead laboratory for two separate studies, gaining AOAC-OMA status for the MBT in December 2017. The first was for confirmation and identification of Salmonella spp., Cronobacter spp., and other Gram-negative organisms (OMA#2017.09). The second was for the confirmation and identification of L. monocytogenes, Listeria spp., and other Gram-positive organisms (OMA#2017.10) in food microbiology.[7, 8] In the meantime, the scope of the AOAC-OMA 2017.09 method has been extended to the confirmation of Campylobacter spp. (awaiting publication). Q Laboratories was also involved in the ISO 16140-6 approvals by MicroVal. Together with the MicroVal technical committee members, the proof of concept was presented with a poster and a symposium at the International Association for Food Protection event in Stockholm in April 2018.[8, 9] Pioneering studies were organized and four certificates were obtained early in 2018.[10]
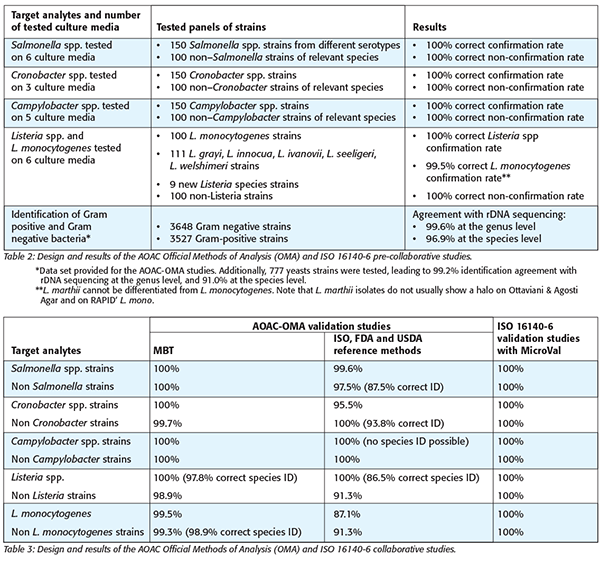 The AOAC-OMA 2017.09 and 2017.10, and the MicroVal Certificates 2017LR72, 2017LR73, 2017LR74, and 2017LR75 cover the confirmation of selected foodborne pathogens and quality indicators, that is, Salmonella spp., Cronobacter spp., Campylobacter spp., Listeria spp., and L. monocytogenes. In addition, the AOAC-OMA 2017.09 and AOAC-OMA 2017.10 include the identification of microbial isolates. The designs and results summary of pre-collaborative and collaborative studies run for the AOAC-OMA and ISO 16140 approvals of the MBT are presented in Tables 2 and 3. The results fit with the requirements of the AOAC Appendix J guidelines and the acceptability limits of the ISO 16140-6 standard.
The AOAC-OMA 2017.09 and 2017.10, and the MicroVal Certificates 2017LR72, 2017LR73, 2017LR74, and 2017LR75 cover the confirmation of selected foodborne pathogens and quality indicators, that is, Salmonella spp., Cronobacter spp., Campylobacter spp., Listeria spp., and L. monocytogenes. In addition, the AOAC-OMA 2017.09 and AOAC-OMA 2017.10 include the identification of microbial isolates. The designs and results summary of pre-collaborative and collaborative studies run for the AOAC-OMA and ISO 16140 approvals of the MBT are presented in Tables 2 and 3. The results fit with the requirements of the AOAC Appendix J guidelines and the acceptability limits of the ISO 16140-6 standard.
Having validation at the level of an AOAC-OMA and ISO validated/MicroVal certified methods is acceptable to the U.S. Food and Drug Administration (FDA) for pathogen confirmation and the European regulations EC 2073/2005.[10] Both FDA and European regulations require a third-party certification body to have approved the validation data and deem that as an equivalent alternative confirmation/identification method to their own methodology, which is considered the gold standard.
Food Microbiology in the Future
MS has historically been an analytical chemistry tool, but the robustness of the MALDI-TOF MS technology in the hands of microbiologists has enabled the Food Microbiology Laboratory to successfully generate spectra and obtain confirmations and identifications within 4 weeks of installing the equipment. Q Laboratories is expanding its identification capabilities and is looking to increase its environmental testing offering, as the need for identification of food and environmental contaminants continues to rise.
As well as at Q Laboratories, there are more than 3,000 MBTs currently running worldwide in various microbiology frameworks. For example:
- Pathogen surveillance at the Pasteur Institute[11] and U.S. Centers for Disease Control and Prevention (CDC) of Atlanta; CDC offers free access to its MicrobeNet library in addition to the Bruker library[12]
- Culture collection curating, such as the Dutch Centraalbureau voor Schimmelcultures, the China Center of Industrial Culture Collection, the Collection de l'Institut Pasteur, the Deutsche Sammlung von Mikroorganismen und Zellkulturen
- For routine testing and quality control purposes
For more information on Q Laboratories, please visit QLaboratories.com.
For more information on Bruker’s MALDI Biotyper, please visit Bruker.com.
- Elbehiry, A, et al. 2017. "Application of MALDI-TOF MS Fingerprinting as a Quick Tool for Identification and Clustering of Foodborne Pathogens Isolated from Food Products." New Microbiologica 40(4):296–278.
- De Koster, C and S Brul. 2016. "MALDI-TOF MS Identification and Tracking of Food Spoilers and Foodborne Pathogens." Curr Opin Food Sci 10:76–84.
- Thouvenot P, Vales G, Bracq-Dieye H, Tessaud-Rita N, M. Maury M, Moura A, Lecuit M, Leclercq A (2018), MALDI-TOF mass spectrometry-based identification of Listeria species in surveillance: A prospective study, Journal of Microbiological Methods Vol 144, pp 29-32.
- Official Methods of Analysis (2016) 20th Ed., AOAC INTERNATIONAL, Rockville, MD, Appendix J. http://www.eoma.aoac.org/app_j.pdf (accessed September 2017)
- ISO 16140-2:2016, TC 34, Microbiology of the food chain—Method validation—Part 2: Protocol for the validation of alternative (proprietary) methods against a reference method.
- ISO/DIS 16140-6:2017, TC 34, Microbiology of the food chain—Method validation—Part 6: Protocol for the validation of alternative (proprietary) methods for microbiological confirmation and typing procedures.
- Bastin, B, et al. 2018. "Confirmation and Identification of Salmonella spp., Cronobacter spp., and Other Gram-Negative Organisms by the Bruker MALDI Biotyper Method: Collaborative Study. First Action 2017.09." J AOAC Int 101(5):1593–1609.
- Bastin, B, et al. 2018. "Confirmation and Identification of Listeria monocytogenes, Listeria spp. and Other Gram-Positive Organisms by the Bruker MALDI Biotyper Method: Collaborative Study, First Action 2017.10." J AOAC Int 101(5):1610–1622.
- Bastin, B, et al. 2018. "Validation of a Confirmation Method According to ISO/DIS 16140-6:2017 A MicroVal Pilot Study using the MALDI Biotyper as an Alternative for Salmonella spp. Confirmation." International Association of Food Protection event, Stockholm, Sweden, Poster P2–16.
- Arbault, P, et al. 2018. "Turning Sequencing and Mass Spectrometry into Routine Testing Tools for Microbial Strain Characterization." International Association of Food Protection event, Stockholm, Sweden, Symposium S19.
- http://microval.org/en/issued-certificates/.
- COMMISSION REGULATION (EC) No 2073/2005 of 15 November 2005 on microbiological criteria for foodstuffs.
- https://www.cdc.gov/microbenet/index.html
Daniele Sohier is the global business development manager, Industrial Microbiology at Bruker Daltonics (Microbiology & Diagnostics business area). With a Ph.D. in biological and medical sciences, Daniele has over 20 years of experience in molecular biology, and has spent the last 15+ years focusing on food microbiology, safety, and quality.


