Foodborne disease remains a substantial burden on public health. The ability to rapidly detect viable pathogens along the production chain is essential for determining intervention and control strategies. Conventional microbiological methods for determining bacterial cell counts employ selective culture, biochemical, and serological characterization. Although these achieve sensitive and selective bacterial detection, they typically require days to weeks for results. To improve the detection speed, the food industry relies on polymerase chain reaction (PCR) for rapid screening of food samples for contamination. Although PCR has gained wide acceptance in the food industry over the past 20 years, the use of expensive equipment and reagents, the need for skilled personnel, and the lack of standardized protocols have hindered its practical implementation for food monitoring and control.[1] In addition, without enrichment, the limit of detection for PCR-based methods is 103–104 CFU/mL.[2] Therefore, it is necessary to associate PCR-based methods with an enrichment step of a few hours.
A large number of biosensor platforms have been developed for rapid on-site, real-time detection of multiple pathogens in food. A biosensor device typically contains a biorecognition element on top of a transducer. The binding of target pathogens with the biorecognition element introduces a signal that can be detected and transmitted by the transducer. A variety of optical, electrochemical, piezoelectric, or thermometric transducers have been employed for biosensing applications. Although many sensor transducers have been reported in the literature, including our own sensor technology,[3–6] applications of biosensors outside research laboratories remain limited. Technical challenges must be addressed, including detection sensitivity (1–3 CFU pathogenic bacteria/25–375 g of a given food product, as defined by the regulatory agencies); matrix effects (to minimize effects from complex food samples); and viable cell detection (should only detect harmful pathogens in food for accurate food safety monitoring).
Preconcentration and Sample Cleanup
Sample preparation methods that remove debris and concentrate pathogens by reducing sample volume are an essential part of the analysis process for detecting small numbers of pathogens.[7,8] Most popular and simple techniques to clean up and concentrate pathogen samples are based on centrifugation[9] and filtration.[10–12] However, significant cell losses can occur during centrifugation, and particulates from the food samples may clog the membrane filters during filtration. Research has been conducted on microfluidics-based cell concentrators that use electrostatic trapping,[13] gravity,[14] surface acoustic wave,[15] dielectrophoresis,[16,17] chemotaxis,[18] and ion-concentration polarization[19] to concentrate cell suspensions. Problems associated with these microfluidic techniques include the limited volume of sample throughput, length of time required to complete the process, and the requirement for relatively clean samples.
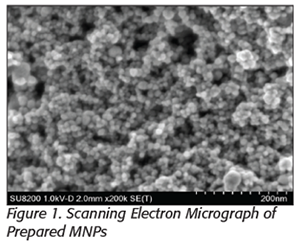 Functionalized magnetic nanoparticles (MNPs) are small magnetic beads (5–100 nm in diameter) that can specifically interact with pathogens. Using cation polymer-coated MNPs, as shown in Figure 1, we have demonstrated a rapid method to recover Gram-negative pathogens from complex food matrices. Bacterial attachment on the nanoparticle is based on electrostatic interactions. After cell capture, an external magnet can be used to recover the attached cells, which can be used in other studies or resuspended in a reduced volume for better detection sensitivity. It has been demonstrated that a greater than 95 percent capture efficacy can be achieved. The attached cells on the MNPs can then be released using a pH 9.25 carbonate buffer. Both capture and preconcentration steps can be performed in 30 minutes, thus reducing the time required for pathogen detection. This novel technology would be useful for a range of industries such as animal processing facilities, vegetable processors, and water purification facilities.
Functionalized magnetic nanoparticles (MNPs) are small magnetic beads (5–100 nm in diameter) that can specifically interact with pathogens. Using cation polymer-coated MNPs, as shown in Figure 1, we have demonstrated a rapid method to recover Gram-negative pathogens from complex food matrices. Bacterial attachment on the nanoparticle is based on electrostatic interactions. After cell capture, an external magnet can be used to recover the attached cells, which can be used in other studies or resuspended in a reduced volume for better detection sensitivity. It has been demonstrated that a greater than 95 percent capture efficacy can be achieved. The attached cells on the MNPs can then be released using a pH 9.25 carbonate buffer. Both capture and preconcentration steps can be performed in 30 minutes, thus reducing the time required for pathogen detection. This novel technology would be useful for a range of industries such as animal processing facilities, vegetable processors, and water purification facilities.
Viable Bacteria Separation via Chemotaxis
Techniques for addressing viability detection problems include the use of ethidium monoazide and propidium monoazide, which have been used to prevent DNA amplification from dead bacterial cells. They specifically permeate dead cells and irreversibly attach to the DNA to prevent amplification.[20–24] However, cost, complexities, and other technical issues have made neither a routine diagnostic tool. Fluorescence microscopy and flow cytometry have also been used to distinguish viable cells from dead cells using a fluorescent dye exclusion method for cell viability measurements. Typical viability dyes are excluded by viable cells but can penetrate membranes of dead cells and emit fluorescence upon binding to nucleic acids inside these cells. However, usage of fluorescent dyes with bacteria is not simple. Due to large interspecies variation, staining and measurement protocols must be developed separately for every bacterial species to ensure qualitative and quantitative high-level results.[25]
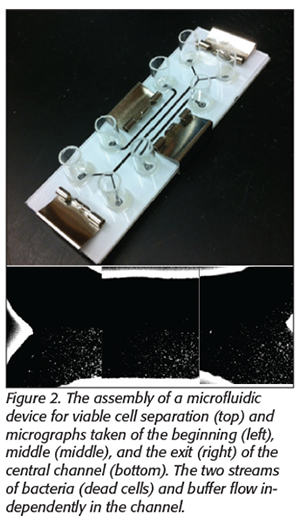 We have explored a novel way to separate viable from dead cells based on bacterial chemotaxis, where bacteria can sense nutrients such as sugars or organic acids and move toward them.[26,27] For example, a microfluidic device with three flow channels, as shown in Figure 2, was fabricated directly on a low-cost nitrocellulose membrane by a laser cutter. The two side channels were used to deliver attractants and repellents for bacterial cells. The center channel had two inlets, one for sample and one for buffer, which converge into a single center channel at the entrance and then split into two collection chambers at the exit with individual downstream collection outlets. The flow inside a microchannel is laminar. Thus, when more than one liquid stream flow next to each other, the only mixing that occurs is by molecular diffusion. The chemoeffectors reach the central channel by diffusing through the nitrocellulose membrane. The dead bacteria stay in the primary sample stream and exit the channel in the collection outlets. However, viable bacteria will sense the chemicals, move toward a preferred location, and leave the channel at a different exit.
We have explored a novel way to separate viable from dead cells based on bacterial chemotaxis, where bacteria can sense nutrients such as sugars or organic acids and move toward them.[26,27] For example, a microfluidic device with three flow channels, as shown in Figure 2, was fabricated directly on a low-cost nitrocellulose membrane by a laser cutter. The two side channels were used to deliver attractants and repellents for bacterial cells. The center channel had two inlets, one for sample and one for buffer, which converge into a single center channel at the entrance and then split into two collection chambers at the exit with individual downstream collection outlets. The flow inside a microchannel is laminar. Thus, when more than one liquid stream flow next to each other, the only mixing that occurs is by molecular diffusion. The chemoeffectors reach the central channel by diffusing through the nitrocellulose membrane. The dead bacteria stay in the primary sample stream and exit the channel in the collection outlets. However, viable bacteria will sense the chemicals, move toward a preferred location, and leave the channel at a different exit.
Initial experiments have shown that the stream of dead bacterial flow does not mix with the buffer stream. Pictures of the beginning, middle, and exit of the center channel are shown in Figure 2, indicating a typical laminar flow in this channel. In contrast, with the application of chemoeffectors (aspartic acid as the attractant and Ni2+ as the repellent) in the proposed design, up to 84 percent of the viable cells made the shift and exited the buffer stream.
Rapid Pathogen Detection
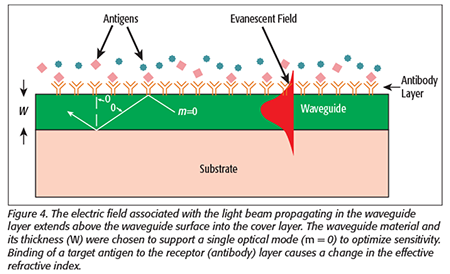
 We have developed a planar waveguide interferometric sensor capable of simultaneously detecting a wide array of analytes in a fast, direct, and sensitive method. The waveguide chip contains an array of sensing interferometers, as shown in Figure 3. Planar waveguides have evanescent fields (the tail of the electromagnetic field associated with the propagation of light), which are sensitive to the index of refraction changes in the volume immediately above the waveguide surface. Coupling a bioreceptor, such as an antibody, to the waveguide surface provides the basis for biological detection (Figure 4). When a test solution containing the target antigen is introduced to the functionalized waveguide, the antigen binds to the receptor and displaces water near the waveguide surface, causing a change in the velocity of the propagating light. To measure this change, an adjacent but unperturbed reference beam is optically combined with the testing beam, creating an interference pattern of alternating dark and light fringes. When refractive index changes occur in the testing arm, the interference pattern shifts, producing a resultant change in the relative phase that is measured using a Fourier transform algorithm and indicating the presence and concentration of the target antigen. No secondary or reporter antibody is required. With detection sensitivities on the order of 0.01 radian, refractive index changes of less than 10-6 can be measured.
We have developed a planar waveguide interferometric sensor capable of simultaneously detecting a wide array of analytes in a fast, direct, and sensitive method. The waveguide chip contains an array of sensing interferometers, as shown in Figure 3. Planar waveguides have evanescent fields (the tail of the electromagnetic field associated with the propagation of light), which are sensitive to the index of refraction changes in the volume immediately above the waveguide surface. Coupling a bioreceptor, such as an antibody, to the waveguide surface provides the basis for biological detection (Figure 4). When a test solution containing the target antigen is introduced to the functionalized waveguide, the antigen binds to the receptor and displaces water near the waveguide surface, causing a change in the velocity of the propagating light. To measure this change, an adjacent but unperturbed reference beam is optically combined with the testing beam, creating an interference pattern of alternating dark and light fringes. When refractive index changes occur in the testing arm, the interference pattern shifts, producing a resultant change in the relative phase that is measured using a Fourier transform algorithm and indicating the presence and concentration of the target antigen. No secondary or reporter antibody is required. With detection sensitivities on the order of 0.01 radian, refractive index changes of less than 10-6 can be measured.
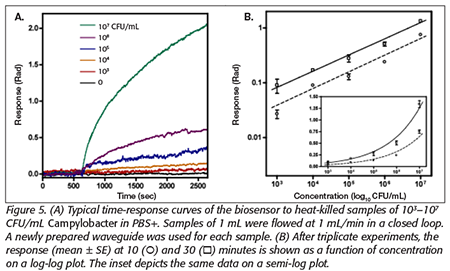
 Current levels of detection are in the femtomolar range for proteins and less than 1,000 CFU/mL for whole organisms. The interferometric sensor provides a direct, near real-time measurement without the need for additional washing or incubation steps. No labeling is required and no consumables other than the buffer solution used for collecting the sample are required. The detection of Campylobacter jejuni (103–107 cells/mL) in PBS using commercially available polyclonal antibody immobilized waveguide is shown in Figure 5. Quantitative reproducibility of the assay was measured using independently prepared sensor chips and found to be within 10 percent over several orders of magnitude in concentration.
Current levels of detection are in the femtomolar range for proteins and less than 1,000 CFU/mL for whole organisms. The interferometric sensor provides a direct, near real-time measurement without the need for additional washing or incubation steps. No labeling is required and no consumables other than the buffer solution used for collecting the sample are required. The detection of Campylobacter jejuni (103–107 cells/mL) in PBS using commercially available polyclonal antibody immobilized waveguide is shown in Figure 5. Quantitative reproducibility of the assay was measured using independently prepared sensor chips and found to be within 10 percent over several orders of magnitude in concentration.
Conclusion
We have witnessed increasing efforts in biosensor development for its great potential for rapid and sensitive pathogen detection. Advances have led to the development of a variety of nanomaterials-based sensor platforms, including metal nanostructure-based surface plasmon resonance, surface-enhanced Raman scattering, carbon nanotube, and graphene-based electrochemical sensors. However, detecting low numbers of viable pathogens in complex matrices is still a hurdle for biosensor application in food safety.
Jie Xu, Ph.D., is a principal research scientist at the Georgia Tech Research Institute.
References
1. Hice, SA, et al. 2019. “Capture, Concentration, and Detection of Salmonella in Foods Using Magnetic Ionic Liquids and Recombinase Polymerase Amplification.” Anal Chem 91(1):1113–1120.
2. Chapela, M-J, A Garrido-Maestu, and AG Cabado. 2015. “Detection of Foodborne Pathogens by qPCR: A Practical Approach for Food Industry Applications.” Cogent Food Agric 1(1):1013771.
3. Schipper, EF, et al. 1998. “The Waveguide Mach-Zender Interferometer as Atrazine Sensor.” Anal Chem 70(6):1192–1197.
4. Schneider, BH, et al. 1997. “Hartman Interferometer: Versatile Integrated Optic Sensor for Label-Free, Real Time Quantification of Nucleic Acids, Proteins, and Pathogens.” Clin Chem 43(9):1757–1763.
5. Xu, J, K Feng, and M Weck. 2011. “Free Chlorine Sensing Using an Interferometric Sensor.” Sens Actuators B: Chemical 156:812–819.
6. Xu, J, D Suarez, and DS Gottfried. 2007. “Detection of Avian Influenza Virus Using an Interferometric Biosensor.” Anal Bioanal Chem 389(4):1193–1199.
7. Ozalp, VC, et al. 2014. “Pathogen Detection by Core-Shell Type Aptamer-Magnetic Preconcentration Coupled to Real-Time PCR.” Anal Biochem 447:119–125.
8. Clime, L, et al. 2015. “Microfluidic Filtration and Extraction of Pathogens from Food Samples by Hydrodynamic Focusing and Inertial Lateral Migration.” Biomed Microdevices 17(1):1–14.
9. Fukushima, H, et al. 2007. “Rapid Separation and Concentration of Food-Borne Pathogens in Food Samples Prior to Quantification by Viable-Cell Counting and Real-Time PCR.” Appl Environ Microbiol 73(1):92–100.
10. Holowecky, PM, et al. 2009. “Evaluation of Ultrafiltration Cartridges for a Water Sampling Apparatus.” J Appl Bacteriol 106(3):738–747.
11. Taku, M. 2012. “Filter-Based Pathogen Enrichment Technology for Detection of Multiple Viable Foodborne Pathogens in 1 Day.” J Food Prot 75(9):1603–1610.
12. Jiang, J, et al. 2014. “Smartphone Based Portable Bacteria Pre-Concentrating Microfluidic Sensor and Impedance Sensing System.” Sens Actuators B 193:653–659.
13. Balasubramanian, AK, et al. 2007. “A Microfluidic Device for Continuous Capture and Concentration of Microorganisms from Potable Water.” Lab Chip 7(10):1315–1321.
14. Warrick, J, et al. 2010. “A Microfluidic Cell Concentrator.” Anal Chem 82(19):8320–8326.
15. Mu, C, et al. 2015. “Development of a Highly Effective Multi-Stage Surface Acoustic Wave SU-8 Microfluidic Concentrator.” Sens Actuators B 215:77–85.
16. Cheng, I-F, T-Y Chen, and H-C Chang. 2014. “Electrokinetics-Based Microfluidic Technology for the Rapid Separation and Concentration of Bacteria/Cells/Biomolecules.” Adv Mat Res 911:347–351.
17. Lewpiriyawong, N, C Yang, and Y Lam. 2012. “Electrokinetically Driven Concentration of Particles and Cells by Dielectrophoresis with DC-Offset AC Electric Field.” Microfluid Nanofluidics 12(5):723–733.
18. Park, S, et al. 2012. “Microfabricated Ratchet Structure Integrated Concentrator Arrays for Synthetic Bacterial Cell-To-Cell Communication Assays.” Lab Chip 12(20):3914–3922.
19. Kwak, R, SJ Kim, and J Han. 2011. “Continuous-Flow Biomolecule and Cell Concentrator by Ion Concentration Polarization.” Anal Chem 83(19):7348–7355.
20. Yang, X, M Badoni, and CO Gill. 2011. “Use of Propidium Monoazide and Quantitative PCR for Differentiation of Viable Escherichia coli from E. coli Killed by Mild or Pasteurizing Heat Treatments.” Food Microbiol 28(8):1478–1482.
21. Melero, B, et al. 2011. “Comparison between Conventional and qPCR Methods for Enumerating Campylobacter jejuni in a Poultry Processing Plant.” Food Microbiol 28(7):1353–1358.
22. Taskin, B, AG Gozen, and M Duran. 2011. “Selective Quantification of Viable Escherichia coli Bacteria in Biosolids by Quantitative PCR with Propidium Monoazide Modification.” Appl Environ Microbiol 77(13):4329–4335.
23. Chen, S, et al. 2011. “Rapid Detection of Viable Salmonellae in Produce by Coupling Propidium Monoazide with Loop-Mediated Isothermal Amplification.” Appl Environ Microbiol 77(12):4008–4016.
24. Liang, N, et al. 2011. “Detection of Viable Salmonella in Lettuce by Propidium Monoazide Real-Time PCR.” J Food Sci 76(4):M234–M237.
25. David, F, et al. 2012. “Viability and Membrane Potential Analysis of Bacillus megaterium Cells by Impedance Flow Cytometry.” Biotechnol Bioeng 109(2):483–492.
26. Celani, A, and M Vergassola. 2010. “Bacterial Strategies for Chemotaxis Response.” Proc Natl Acad Sci U.S.A. 107(4):1391–1396.
27. Miller, LD, et al. Chapter 3, “Diversity in Bacterial Chemotactic Responses and Niche Adaptation.” In Advances in Applied Microbiology (Academic Press, 2009), 53–75.
Biosensor-Based Rapid Pathogen Detection: Are We There Yet?