The food safety industry has remained assiduous in developing methods and materials that test for the presence of bacterial contamination. Despite this diligence, circumstances occasionally arise for which there are no ready answers. Test method developers and test kit manufacturers must address the most frequently occurring problems first, in order to make their test kits commercially viable. Proprietary test kits are manufactured to address the most common bacteria, such as E. coli, Campylobacter, Listeria, and Salmonella, but what if an unfamiliar bacterium is the cause of the adulteration? When a problem arises for which there is no test kit available, a qualified contract laboratory can design a protocol specifically for the bacterial dilemma utilizing polymerase chain reaction (PCR) technology. In essence, the lab can custom design a test kit that addresses the specific predicament that is encountered.
Preventing microbiological contamination in food production facilities, and subsequently in food products, is an ongoing and frequently challenging proposition. Having the best methods available to measure the cleaning procedures, the purity of incoming raw materials, the quality of a process or product, or the presence of bacterial contamination is paramount to maintaining the quality of a product. With an efficient and effective sampling plan in place and with all of the latest testing methods available, a food company should be problem free, right? Not necessarily. Many bacteria present difficult challenges, especially when the contaminant, although not necessarily pathogenic, negatively affects the quality of the product. Many times in this scenario, the frustration is exacerbated because there is no test kit or available method to screen for the offending bacterium.
Finding Answers
Let’s take a look at a possible scenario of bacterial contamination for which a custom test kit may need to be designed. A product has become adulterated by a bacterium such as a Lactobacillus spp., Acetobacter spp., or any number of other bacteria for which there are no “species specific” test kits currently available. Both Lactobacillus and Acetobacter have several species in their respective genus that may occur in a facility. Each may appear in a product from time to time, regardless of how strictly adhered to the sanitation schedule or how thoroughly a Hazard Analysis & Critical Control Points (HACCP) plan is applied. Presence of these organisms in a manufacturing or processing environment may not have caused much concern in the past, but then something changes. Suddenly the product—be it raw beef an artificial coloring, or a flavor substitute— starts to develop problems. The color may be off, the taste may have changed, the product may be reacting and breaking down into other undesired components. The problem needs to be addressed in the most expeditious and effective means possible.
After poring over previous test results, it is discovered that the problem has occurred before, though only sporadically. Now it is appearing with greater frequency. The data from previous tests indicates that each time this problem has arisen, there have been high Lactobacillus counts. Testing from those lots indicated a particular Lactobacillus spp. was identified each time the problem occurred, growing in concert with several other Lactobacillus species. This specific Lactobacillus spp. metabolizes in such a way that it affects the chemistry of the product, causing the deleterious effect. This problem bacterium could easily be a Lactobacillus species that occasionally shows itself in the facility, or even one that has mutated recently in the facility, changing its metabolism in such a way that what used to be just another spoilage organism now causes major issues with product quality. The methods currently available are not specific enough to identify this particular problem bacterium. The compendial test for Lactobacillus is a routine plating method, only useful for getting a general idea of the total microbial load of Lactobacillus spp. in a product. It is especially frustrating to correctly identify the bacteria, but discover there is no accurate way to test for it with a commercial kit, because none is available.
Customizing Solutions
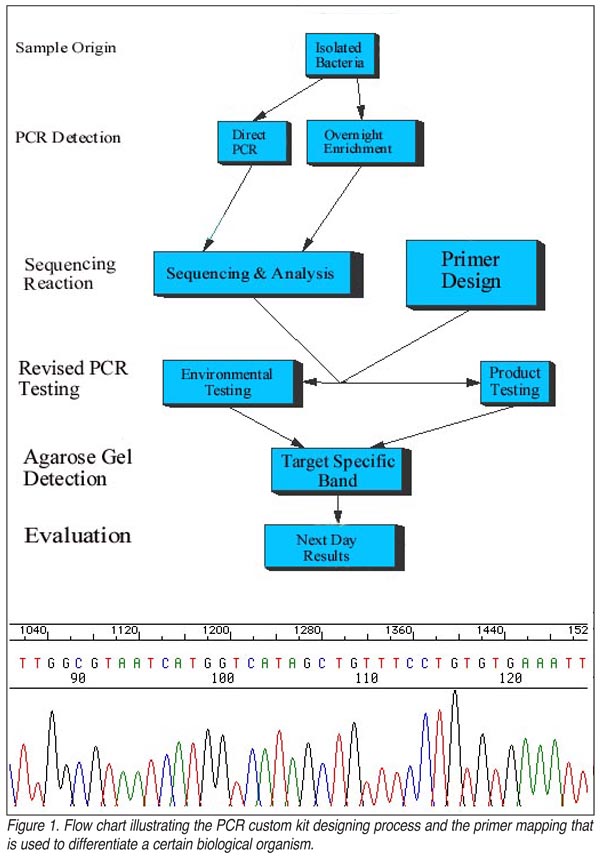 It is possible to custom design a kit and screen for a specific bacterial species by using PCR technology (Figure 1). This can be used on a product sample, a swab, a sponge or other sampling method, and can be utilized the day following sampling, after a brief overnight enrichment. This will replace current methods involving traditional microbiological techniques. PCR detects bacteria through identifying and amplifying a segment of DNA specific for a particular organism. Since certain portions of DNA are unique for every living thing, we can use this distinctiveness as a means of detection. Testing by PCR will increase the effectiveness of a HACCP plan, as well as aid in effectively identifying any areas contaminated with the problem bacterium. In addition to the unparalleled accuracy of PCR analysis, results are usually available within 24 to 48 hours. Obtaining results quickly allows identification of probable areas and conduits of contamination, enabling a reevaluation of the sanitation plan and an increased ability to make adjustments where needed.
It is possible to custom design a kit and screen for a specific bacterial species by using PCR technology (Figure 1). This can be used on a product sample, a swab, a sponge or other sampling method, and can be utilized the day following sampling, after a brief overnight enrichment. This will replace current methods involving traditional microbiological techniques. PCR detects bacteria through identifying and amplifying a segment of DNA specific for a particular organism. Since certain portions of DNA are unique for every living thing, we can use this distinctiveness as a means of detection. Testing by PCR will increase the effectiveness of a HACCP plan, as well as aid in effectively identifying any areas contaminated with the problem bacterium. In addition to the unparalleled accuracy of PCR analysis, results are usually available within 24 to 48 hours. Obtaining results quickly allows identification of probable areas and conduits of contamination, enabling a reevaluation of the sanitation plan and an increased ability to make adjustments where needed.
After isolating and identifying the bacterial species, it is possible for a reference lab to design a specialized PCR detection assay. This initial set-up may take a few days to implement, but once the work is done the result will be a rapid method specific to the problem. Designing a specific PCR assay involves three steps: (1) sequencing and primer design; (2) enrichment determination, minimum threshold level of detection and optimization; and (3) implementing a testing program and applying the results.
Step 1: Sequencing and Primer Design. The first step in designing a kit is to find out what makes the problem bacteria unique from other bacteria. This is accomplished by looking at the basic building blocks that make every living thing on earth unique—the DNA. A reference lab can submit the bacteria for sequencing, and once they obtain the sequence, they can compare it to other sequences of closely related bacteria and search for differences between them. These differences are what will be used to detect the problem bacteria from all other bacteria in the sample. This is how primers, which are short pieces of DNA that sit down on target DNA and facilitate replication, are designed.
Specifically, the lab will amplify some relatively conserved region of its genome (the 16S rDNA gene is popular) using degenerate primers, sequence it and design new, organism-specific primers. It is this ability to concentrate on the DNA of the organism in question that will enable the laboratory to construct a testing protocol that will screen exclusively for the particular species, thereby gaining the advantage over traditional methods that may only specify the presence of a particular genus.
Step 2: Enrichment and PCR Optimization. The second step is to increase the “signal-to-noise” ratio of the PCR. In other words, the protocol must be crafted in such a way so as to detect exclusively for the species desired (“the signal”) and not complicate or confuse the results by incorrectly detecting for other bacteria (“noise”). The optimal point in the process to increase the signal and decrease the noise is the enrichment step. The samples will contain many other types of bacteria, some similar to the target, some not so similar. The enrichment step typically is an eight- to 24-hour incubation period in a broth designed to grow the target bacteria and recover damaged cells, while eliminating as many other types of bacteria as possible, many of which may interfere or hinder the efficiency of the PCR reaction. This takes a bit of research to discover what additives, sugars and other components can be added to enhance the growth of the targeted bacteria, as well as what chemical or antibiotic agents minimize the growth of other bacteria that may otherwise out- compete the target bacteria.
The purpose of this critical step is to encourage the elimination of cells that are not the target while simultaneously encouraging the target cells (which may be weak and damaged from processing) to grow. Another common problem in any assay for microorganisms, especially in processed foods, is the recovery of damaged organisms. The PCR process has been optimized to amplify the DNA of only viable organisms, thus eliminating the possibility of false-positive results.
Step 3: Implementation of Testing Program. Once the test kit is designed, it can provide useful and meaningful data in a very short period of time. One of the most useful types of information this testing program can supply is data that helps the manufacturer determine where to sample in the plant. The types of samples to take are always problem-dependent, and therefore, a review of the specific problem should dictate what samples are taken. For example, custom PCR test data may indicate that a plant should sample finished products only, or that sampling everything from raw materials to every piece of equipment on the production line is the most effective course of action for a particular food manufacturing facility. Data may show that this could be done with swabs or sponges, or by testing product samples directly, depending on the nature of the problem.
The optimal program is to combine both sample analysis and environmental testing to isolate the cause of the contamination and to undertake measures to eradicate the problem. Samples should be taken from all suspected areas of contamination, and sampling can be scaled back as potential problem areas are eliminated. The advantages of PCR technology are speed and specificity, enabling the food company to implement corrective measures very quickly. If positives come up for the target bacteria, those areas should be sanitized immediately and sampled again. If the results yield positives in many areas that do not indicate an initial point of entry, the scope of sampling must be reevaluated and restructured to cover a wider area. If the target is found in a raw material used in production, the solution may be to begin screening the incoming raw material prior to use. If the method of production, the facility or any number of other things change, the sampling scheme and HACCP plan should change along with it. The results from this PCR assay need to be analyzed and applied effectively to ensure that the cause of contamination has been located and eliminated.
Conclusion
Despite the continued efforts of manufacturers and regulatory bodies to develop testing methods and materials to screen for food contaminants, research and development has primarily focused on the most frequently occurring bacteria. A food production facility is an evolving, ever-changing ecosystem with myriad potentially harmful bacteria. When a bacterium invades a product or production location, there is not time to wait for R&D companies to develop and market a test kit for the particular bacterium that is wreaking havoc. Instead, it may be prudent to work with the testing laboratory to develop a protocol using PCR to screen specifically for the organism instigating the contamination.
John R. Lewis, Jr., M.S., is Microbiology Group Leader for Q Laboratories, Inc, an independent microbiology and chemistry analytical laboratory located in Cincinnati, OH. He holds a master’s degree in molecular biology and has been utilizing PCR technology for more than a decade. Prior to coming to Q Laboratories, where he has worked for nearly four years, Lewis worked at the nationally recognized Cincinnati Children’s Hospital performing cancer research using PCR and other molecular techniques. Lewis can be reached at jlewis@qlaboratories.com.
Using PCR to Custom Design a Test Kit